Spatio-Temporal AI & Graph Learning: Some papers that i went through
Introduction
If you’re diving into research on biological systems, dynamic networks, or AI that learns from complex, time-evolving data, you’ve probably stumbled into a overwhelming jungle of papers. As I’m building my GSOC 2026 proposal for the Openworm Foundation, I realized there are some papers that fundamentally shift how we think about AI and biological systems.
Here’s the reality: most biological systems are not static. Brains evolve, cells communicate, organisms adapt. Yet many traditional AI models treat the world like a frozen snapshot. These five papers challenge that assumption—and in doing so, they’re reshaping the entire landscape of spatio-temporal AI and graph learning.
Let me break down what these papers do, why they matter, and why you should care about them.
1. ESTGEL: Understanding How Brain Networks Evolve [1]
Paper: Explainable Spatio-Temporal Graph Evolution Learning with Applications to Dynamic Brain Network Analysis During Development
The Big Picture
Imagine your brain as a living, breathing network of connections. It’s not the same today as it was when you were 10—and it won’t be the same at 60. ESTGEL captures exactly this: how brain networks structurally and functionally reorganize as we grow.
How It Works
Instead of traditional approaches that treat each time step independently, ESTGEL uses:
- Edge attention modules: Focuses on which connections matter most at each moment
- Dynamic gating mechanisms: Remembers past states and predicts future ones
- Explainability built-in: You can actually understand why the model made a decision
The key insight? By analyzing functional MRI (fMRI) data from developing brains, ESTGEL reveals that as we age, our brain networks transition from highly dispersed to concentrated, efficient structures. It’s like watching the internet evolve—from a chaotic mess to an optimized highway system.
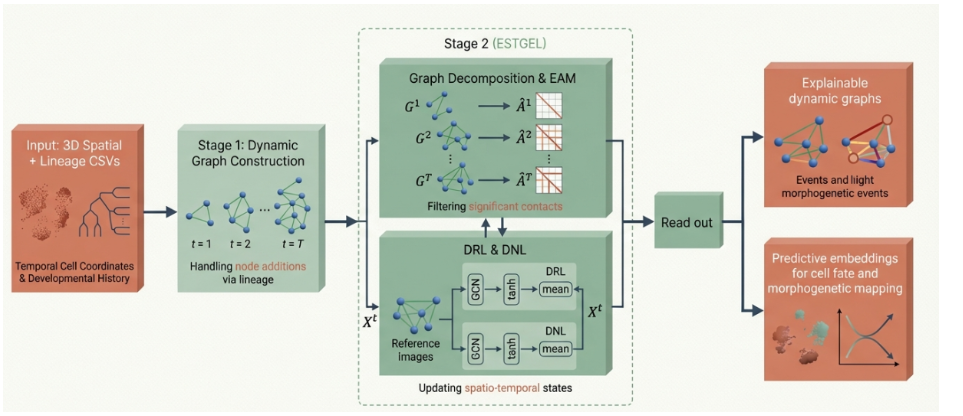 ESTGEL model architecture for analyzing dynamic brain network evolution.
ESTGEL model architecture for analyzing dynamic brain network evolution.
From a developmental biology perspective, this is huge. For organisms like C. elegans, we have the complete connectome mapped, but we don’t fully understand how it changes over development. ESTGEL’s approach—combining graph attention with temporal dynamics—could unlock similar insights in simpler organisms.
2. ESA: The Beauty of Simplicity in Graph AI [2]
Paper: An End-to-End Attention-Based Approach for Learning on Graphs (Edge Set Attention)
Graph Neural Networks (GNNs) are notorious for their complexity: positional encodings, message-passing schemes, pre-processing pipelines… it’s a lot. ESA throws most of that away and proves that sometimes, simplicity wins.
How It Works
Edge Set Attention (ESA) takes a radical approach:
- Treat a graph as a set of edges (connections between nodes)
- Use pure attention mechanisms (like ChatGPT uses)
- Alternate between:
- Masked attention: Only look at directly connected edges
- Self-attention: See the bigger picture
That’s it. No complex pre-processing. No positional encodings. Yet…
Despite being explicitly designed for simplicity, ESA outperforms heavily fine-tuned complex graph transformers across dozens of benchmarks:
- Molecular property prediction
- Vision tasks
- Classification problems
This is the kind of result that makes you question everything. It proves that architectural elegance often defeats brute-force complexity.
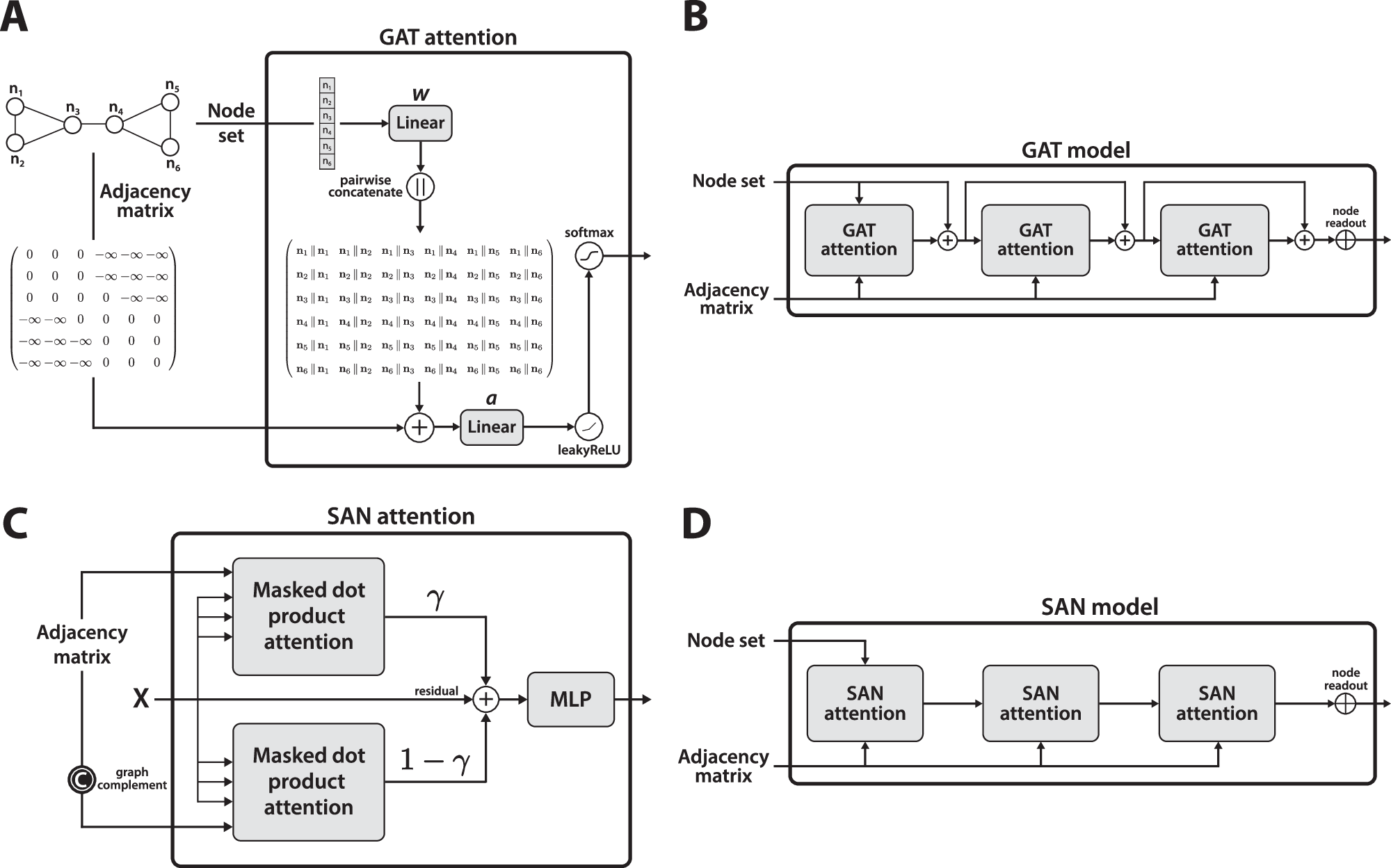 ESA’s end-to-end attention mechanism for graph learning—simple yet powerful.
ESA’s end-to-end attention mechanism for graph learning—simple yet powerful.
For biological systems (proteins, metabolic networks, gene interactions), we often need to represent them as graphs. ESA’s simplicity and effectiveness make it an ideal candidate for tasks like predicting protein-protein interactions or understanding metabolic pathways.
3. Self-Referential GHNs: AI That Evolves Itself [3]
Paper: Hypernetworks That Evolve Themselves
Here’s a mind-bending idea: What if your AI model could literally mutate itself?
In nature, organisms don’t get a PhD in optimization—they evolve through random mutation and environmental selection. This paper asks: why can’t neural networks do the same?
How It Works
A Self-Referential Graph HyperNetwork (GHN) embeds evolution directly into its own code:
- The network looks at its own structure
- Generates mutated versions of itself
- Tests them in the environment
- Keeps the winners, discards the losers
No external algorithm tweaking weights. The network becomes the optimizer.
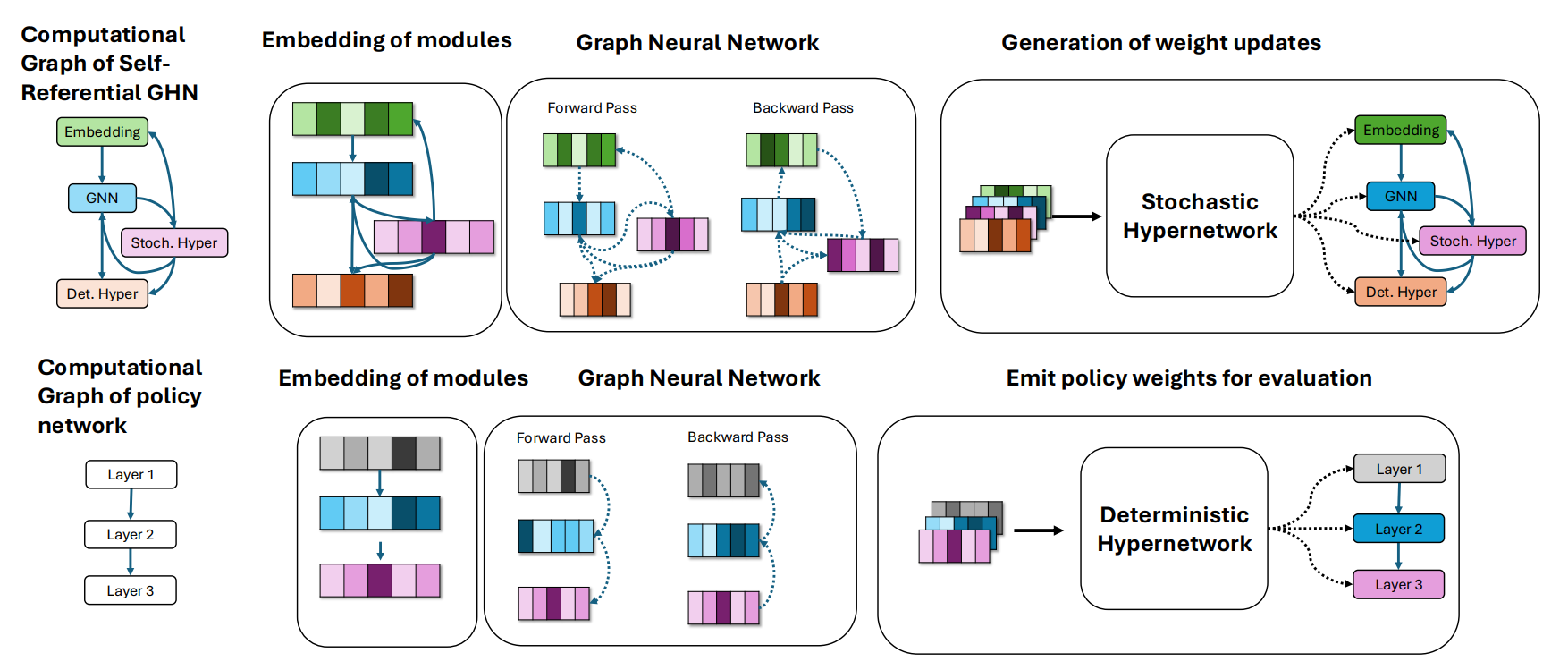 HyperNetwork architecture enabling self-referential evolution.
HyperNetwork architecture enabling self-referential evolution.
Here’s where it gets wild: the network exhibits emergent control over its own evolvability:
- Environment stable? → Reduce mutation rate (fine-tune performance)
- Environment changing? → Crank up mutation rate (explore new solutions)
- Novel challenge? → The network figures out it needs to change strategies
This is the first step toward truly autonomous, open-ended learning agents—AI that doesn’t need a human telling it what to do.
For developmental biology, this is conceptually similar to how organisms adapt during development. Neural plasticity, for example, is the brain’s ability to rewire itself based on experience. If we can achieve something similar in AI, we’re not just mimicking nature—we’re understanding it at a deeper level.
4. Cell-STN: The Ultimate Cell Tracker [4]
Paper: Spatio-Temporal Feature Based Deep Neural Network for Cell Lineage Analysis in Microscopy Images
Biologists spend hours—sometimes days—manually tracking cells under a microscope:
- Where did this cell move?
- When did it divide (mitosis)?
- When did it die (apoptosis)?
Cell-STN automates this nightmare.
How It Works
Cells aren’t static objects; they change shape, move, and behave unpredictably. Standard computer vision isn’t enough. Cell-STN uses:
- Bidirectional ConvLSTMs: Scan video frames both forward and backward in time
- Multi-task learning: Predict locations, divisions, and deaths simultaneously
- Visual + temporal features: Learn both what cells look like AND how they move
The result? AI that understands cell behavior in a holistic way.
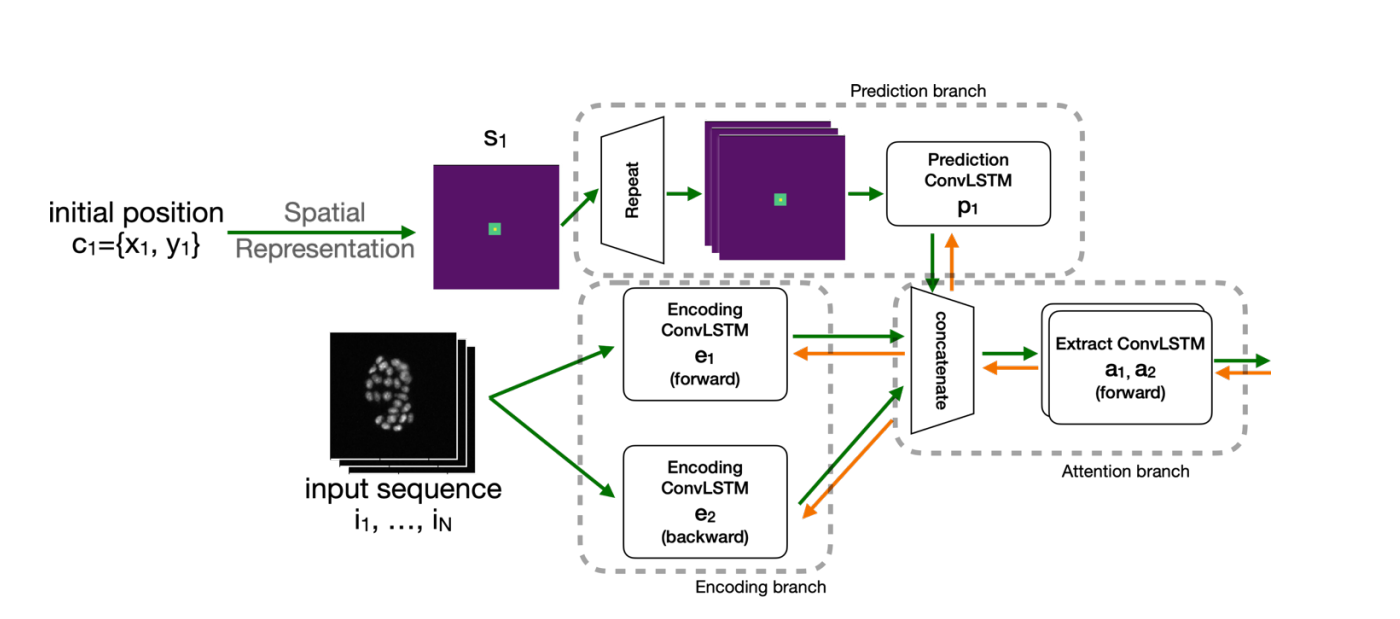 Cell-STN bidirectional ConvLSTM architecture for temporal cell tracking.
Cell-STN bidirectional ConvLSTM architecture for temporal cell tracking.
Here’s what makes this special: it only requires weakly supervised, coarse labels. No need to perfectly trace every cell outline. This means researchers can use it on massive video datasets without spending their entire PhD on manual annotation.
For Developmental Biology
This is perfect for studying organisms like C. elegans. We could use Cell-STN to:
- Track cell divisions during development
- Understand cell migration patterns
- Study developmental abnormalities
It bridges the gap between microscopy data and computational biology.
5. CellNEST: Decoding Cellular “Gossip” [5]
Paper: CellNEST Reveals Cell–Cell Relay Networks Using Attention Mechanisms on Spatial Transcriptomics
Cells communicate through a language of proteins (ligands) binding to receptors. Existing tools catch individual conversations, but they miss the plot: cells can relay messages to other cells, creating networks of communication.
CellNEST decodes these relay networks.
How It Works
Using Graph Attention Networks (GAT) trained in an unsupervised manner, CellNEST:
- Analyzes spatial transcriptomic data
- Identifies which cells communicate with which
- Detects relay patterns (A → B → C)
- Outputs high-confidence communication maps
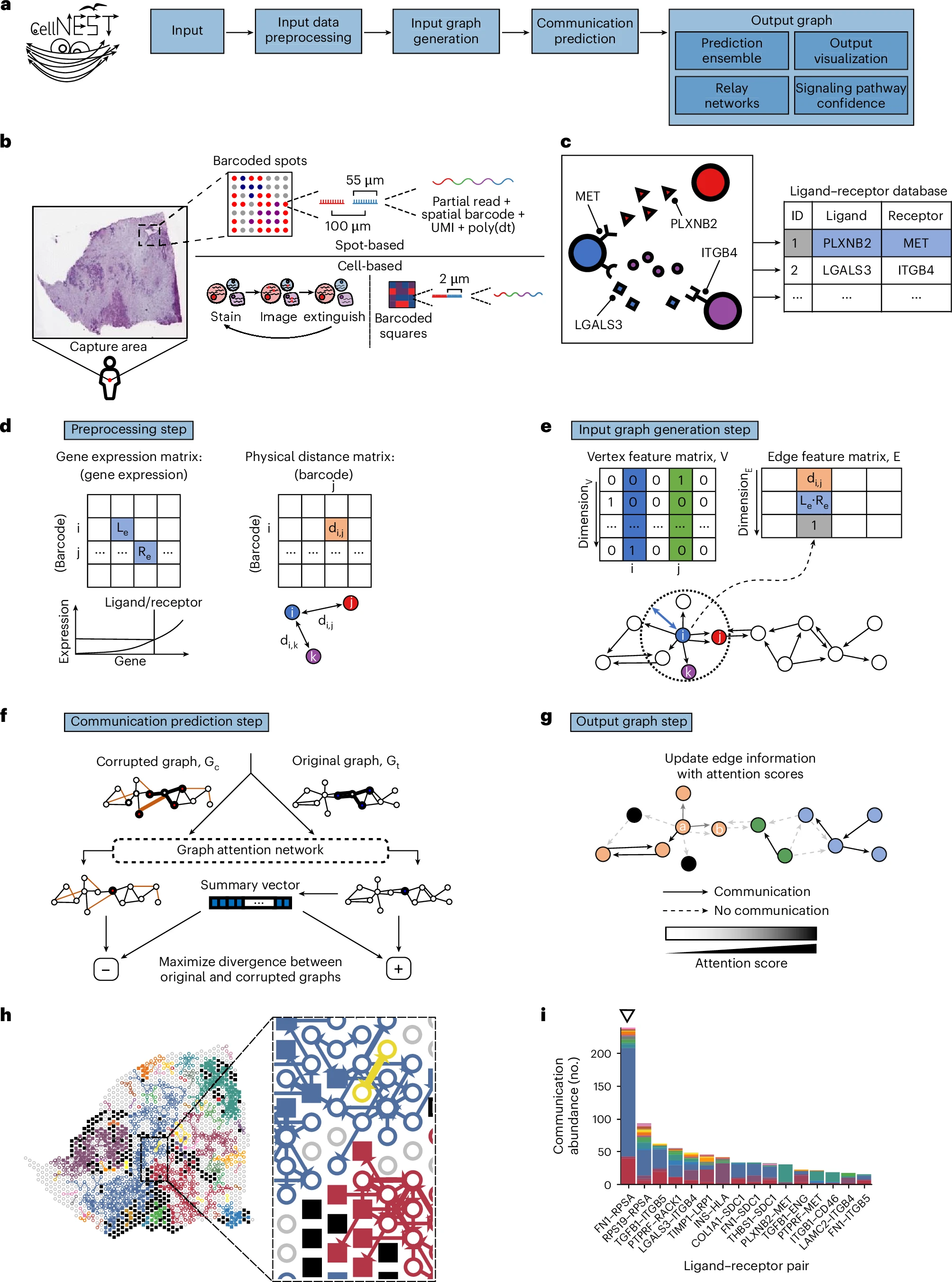 CellNEST reveals relay networks and cell-cell communication patterns in tissue.
CellNEST reveals relay networks and cell-cell communication patterns in tissue.
Real-World Impact
CellNEST successfully mapped:
- T cell homing signals in lymph nodes
- Aggressive communication networks in pancreatic and colorectal tumors
For the first time, researchers could visualize not just who’s talking to whom, but how information cascades through tissues.
Cell-cell communication is the foundation of development, immunity, and disease. If we truly understand these relay networks, we can:
- Predict how diseases spread
- Design better immunotherapies
- Understand developmental signaling cascades
What Connects All Five Papers?
If I had to find the throughline, it’s this:
Modern AI for biology isn’t about finding static patterns; it’s about understanding dynamic, interconnected systems.
- ESTGEL shows brain networks evolve
- ESA proves simplicity can beat complexity
- Self-Referential GHNs suggest AI can adapt on its own
- Cell-STN automates tracking over time
- CellNEST reveals hidden communication networks
All five treat their respective domains (brains, graphs, AI itself, cells, tissues) as living, changing systems—not frozen snapshots.
Implications for My GSOC Work
As I work on my proposal for the Openworm Foundation, these papers are reshaping my thinking:
- Graph attention mechanisms (ESA, CellNEST) are probably the right tool for understanding C. elegans connectomics
- Spatio-temporal models (ESTGEL, Cell-STN) could help us understand how the connectome changes during development
- Self-referential learning (Self-Ref GHNs) hints at how neural systems might optimize themselves—something we could study in C. elegans
The convergence is clear: the future of computational biology is about dynamic graphs, attention mechanisms, and systems that learn to evolve.
Final Thoughts
When I first dove into these papers, I thought I was just gathering random research. But the more I read, the more I realized they’re not random at all. They’re all pieces of a larger puzzle: how do we build AI that understands biology as it actually is—dynamic, interconnected, and evolving?
If you’re working on anything related to:
- Neural networks
- Cell biology
- Developmental biology
- Graph learning
- Biological simulations
…you need to know these papers.
So yeah, I’m excited about my GSOC proposal. These five papers just gave me a great insight!